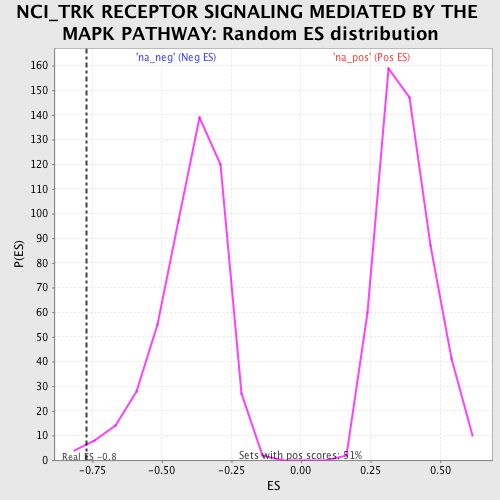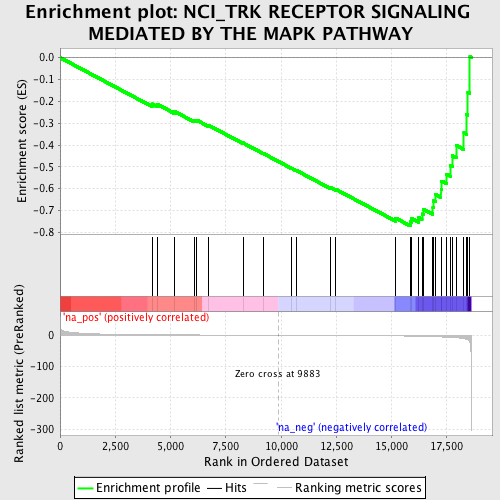
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_DMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | NCI_TRK RECEPTOR SIGNALING MEDIATED BY THE MAPK PATHWAY |
| Enrichment Score (ES) | -0.770507 |
| Normalized Enrichment Score (NES) | -1.9308457 |
| Nominal p-value | 0.008097166 |
| FDR q-value | 0.7058529 |
| FWER p-Value | 0.873 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RAP1A | 4163 | 1.660 | -0.2120 | No | ||
| 2 | MAPK7 | 4405 | 1.547 | -0.2139 | No | ||
| 3 | RIT2 | 5180 | 1.261 | -0.2464 | No | ||
| 4 | NTF3 | 6066 | 0.983 | -0.2870 | No | ||
| 5 | HRAS | 6193 | 0.946 | -0.2869 | No | ||
| 6 | MAP2K5 | 6735 | 0.790 | -0.3103 | No | ||
| 7 | CDK5R1 | 8277 | 0.392 | -0.3904 | No | ||
| 8 | NRAS | 9203 | 0.170 | -0.4390 | No | ||
| 9 | FOS | 10490 | -0.151 | -0.5071 | No | ||
| 10 | RPS6KA1 | 10686 | -0.204 | -0.5161 | No | ||
| 11 | SRF | 12241 | -0.633 | -0.5951 | No | ||
| 12 | MAP2K1 | 12474 | -0.704 | -0.6026 | No | ||
| 13 | RUSC1 | 15180 | -1.932 | -0.7342 | No | ||
| 14 | MAP2K3 | 15856 | -2.564 | -0.7520 | Yes | ||
| 15 | MAP3K2 | 15889 | -2.600 | -0.7350 | Yes | ||
| 16 | RAF1 | 16233 | -3.067 | -0.7314 | Yes | ||
| 17 | PRKCD | 16381 | -3.304 | -0.7155 | Yes | ||
| 18 | CDK5 | 16438 | -3.438 | -0.6938 | Yes | ||
| 19 | MAPK14 | 16874 | -4.333 | -0.6860 | Yes | ||
| 20 | CREB1 | 16910 | -4.443 | -0.6559 | Yes | ||
| 21 | MAP2K6 | 16991 | -4.589 | -0.6271 | Yes | ||
| 22 | RAP1B | 17239 | -5.306 | -0.6022 | Yes | ||
| 23 | MAPK1 | 17264 | -5.380 | -0.5647 | Yes | ||
| 24 | MAPK3 | 17495 | -6.022 | -0.5337 | Yes | ||
| 25 | RPS6KA5 | 17680 | -6.679 | -0.4955 | Yes | ||
| 26 | RIT1 | 17758 | -6.945 | -0.4497 | Yes | ||
| 27 | MAPKAPK2 | 17938 | -7.889 | -0.4025 | Yes | ||
| 28 | EHD4 | 18277 | -10.570 | -0.3446 | Yes | ||
| 29 | KRAS | 18389 | -12.525 | -0.2603 | Yes | ||
| 30 | MEF2C | 18457 | -14.583 | -0.1589 | Yes | ||
| 31 | EGR1 | 18550 | -23.249 | 0.0036 | Yes |